Hannover Medical School researcher Prof. Galardini from the RESIST Cluster of Excellence has found causes for bloodstream infections in the genes of bacteria. This will enable better diagnostics and vaccinations in the future.
Escherichia coli bacteria live in the intestines of humans and play an important role there for normal intestinal function as well as for a functioning immune system. These intestinal inhabitants do not form a uniform population, but consist of a large number of strains that differ greatly in their genome and also in their metabolism.
Most strains of E. coli are harmless, but some can cause diarrhea or urinary tract infections and—if they enter the blood—bloodstream infections and sepsis via their toxins. Sepsis is the third most common cause of death in Germany.
The bacteria have caused increasingly more diseases
Prof. Dr. Marco Galardini’s team has found that E. coli has a significant genetic variation that contributes to the transition between the harmless life in the intestine (commensalism) and the pathogenic form. In addition, the researchers were also able to show that this bacterial species has evolved more toward causing disease over the years.
“Building on these findings, we envision the creation of better molecular diagnostic tools in the future, and these results might also be important for vaccines development,” says Prof. Galardini.
The team published their findings in the journal PLoS Genetics. First authors are Judit Burgaya and Julie Marin. The work originated in TWINCORE in collaboration with Prof. Erick Denamur (INSERM, Paris) and Prof. François Blanquart (Collège de France).
Significant genetic differences between harmless and dangerous bacteria
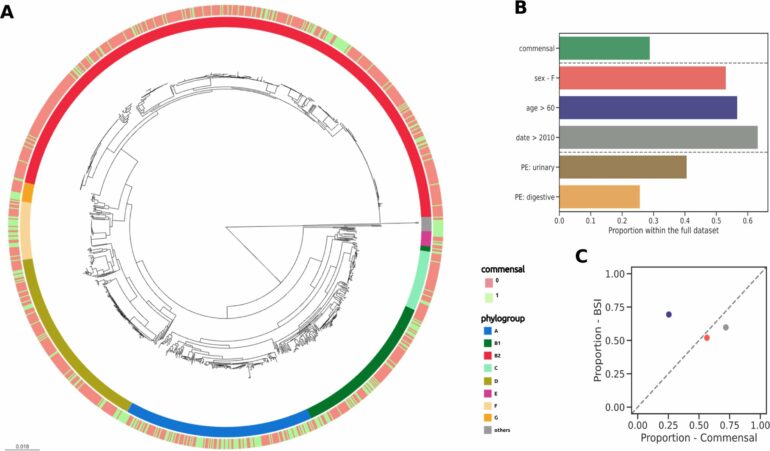
The team examined a collection of about 900 E. coli isolates that caused blood infections and 370 harmless isolates. The samples were collected over a 17-year period (from 2000 to 2017) by the team of Prof. Erick Denamur.
“We found significant differences between the disease-causing and the harmless isolates—both in their pangenomes, i.e. the totality of genes of the respective isolates, and in their genetic backgrounds, in terms of the presence of virulence-associated genes and antimicrobial resistance genes,” says Prof. Galardini.
Using another commensal collection from 1980, the group also found that pathogenicity might have increased steadily from 1980 through 2000 to 2010.
This work is the third in a series of studies aimed at understanding the genetic determinants of the ability of E. coli to cause bloodstream infections. The team published the first two papers in 2020 and 2022.
More information:
Judit Burgaya et al, The bacterial genetic determinants of Escherichia coli capacity to cause bloodstream infections in humans, PLOS Genetics (2023). DOI: 10.1371/journal.pgen.1010842
Provided by
Medizinische Hochschule Hannover
Citation:
How harmless E. coli turns dangerous (2023, August 18)