Artificial intelligence has changed the way science is done by allowing researchers to analyze the massive amounts of data modern scientific instruments generate. It can find a needle in a million haystacks of information and, using deep learning, it can learn from the data itself. AI is accelerating advances in gene hunting, medicine, drug design and the creation of organic compounds.
Deep learning uses algorithms, often neural networks that are trained on large amounts of data, to extract information from new data. It is very different from traditional computing with its step-by-step instructions. Rather, it learns from data. Deep learning is far less transparent than traditional computer programming, leaving important questions – what has the system learned, what does it know?
As a chemistry professor I like to design tests that have at least one difficult question that stretches the students’ knowledge to establish whether they can combine different ideas and synthesize new ideas and concepts. We have devised such a question for the poster child of AI advocates, AlphaFold, which has solved the protein-folding problem.
Protein folding
Proteins are present in all living organisms. They provide the cells with structure, catalyze reactions, transport small molecules, digest food and do much more. They are made up of long chains of amino acids like beads on a string. But for a protein to do its job in the cell, it must twist and bend into a complex three-dimensional structure, a process called protein folding. Misfolded proteins can lead to disease.
In his chemistry Nobel acceptance speech in 1972, Christiaan Anfinsen postulated that it should be possible to calculate the three-dimensional structure of a protein from the sequence of its building blocks, the amino acids.
Just as the order and spacing of the letters in this article give it sense and message, so the order of the amino acids determines the protein’s identity and shape, which results in its function.
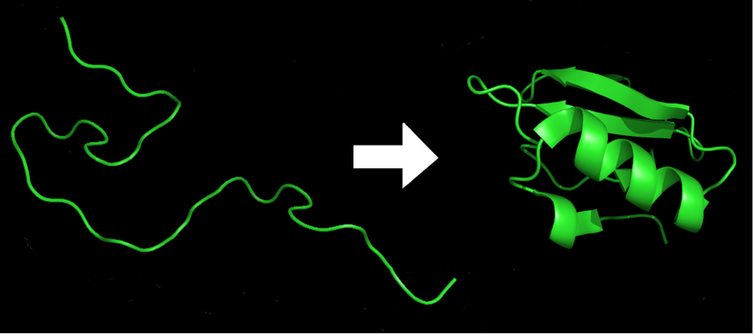
Within milliseconds of the exit of an amino acid chain (left) from the ribosome, it is folded into the lowest-energy 3D shape (right), which is required for the protein’s function.
Marc Zimmer, CC BY-ND
Because of the inherent flexibility of the amino acid building blocks, a typical protein can adopt an estimated 10 to the power of 300 different forms. This is a massive number, more than the number of atoms in the universe. Yet within a millisecond every protein in an organism will fold into its very own specific shape – the lowest-energy arrangement of all the chemical bonds that make up the protein. Change just one amino acid in the hundreds of amino acids typically found in a protein and it may misfold and no longer work.
AlphaFold
For 50 years computer scientists have tried to solve the protein-folding problem – with little success. Then in 2016 DeepMind, an AI subsidiary of Google parent Alphabet, initiated its