Neural networks (NNs) are increasingly being used to predict new materials, the rate and yield of chemical reactions, and drug-target interactions, among others. For these applications, they are orders of magnitude faster than traditional methods such as quantum mechanical simulations.
The price for this agility, however, is reliability. Because machine learning models only interpolate, they may fail when used outside the domain of training data.
But the part that worried Rafael Gómez-Bombarelli, the Jeffrey Cheah Career Development Professor in the MIT Department of Materials Science and Engineering, and graduate students Daniel Schwalbe-Koda and Aik Rui Tan was that establishing the limits of these machine learning (ML) models is tedious and labor-intensive.
This is particularly true for predicting ”potential energy surfaces” (PES), or the map of a molecule’s energy in all its configurations. These surfaces encode the complexities of a molecule into flatlands, valleys, peaks, troughs, and ravines. The most stable configurations of a system are usually in the deep pits—quantum mechanical chasms from which atoms and molecules typically do not escape.
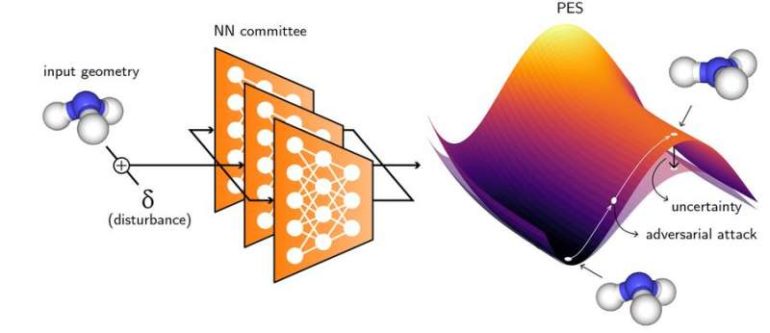
In a recent Nature Communications paper, the research team presented a way to demarcate the “safe zone” of a neural network by using “adversarial attacks.” Adversarial attacks have been studied for other classes of problems, such as image classification, but this is the first time that they are being used to sample molecular geometries in a PES.
“People have been using uncertainty for active learning for years in ML potentials. The key difference is that they need to run the full ML simulation and evaluate if the NN was reliable, and if it wasn’t, acquire more data, retrain and re-simulate. Meaning that it takes a long time to nail down the right model, and one has to run the ML simulation many times” explains Gómez-Bombarelli.
The Gómez-Bombarelli lab at MIT works on a synergistic synthesis of first-principles simulation and machine learning that greatly speeds up this process. The actual simulations are run only for a small fraction of these molecules, and all those data are fed into a neural network that learns how to predict the same properties for the rest of the molecules. They have successfully demonstrated these methods for a growing class of novel materials that includes catalysts for producing hydrogen from water, cheaper polymer electrolytes for electric vehicles, zeolites for molecular sieving, magnetic materials, and more.
The challenge, however, is that these neural networks are only as smart as the data they are trained on. Considering the PES map, 99 percent of the data may fall into one pit, totally missing valleys that are of more interest.
Such wrong predictions can have disastrous consequences—think of a self-driving car that fails to identify a person crossing the street.
One way to find out the uncertainty of a model is to run the same data through multiple versions of it.
For this project, the researchers had multiple neural networks predict the potential energy surface from the same data. Where the network is fairly sure of the prediction, the variation between the outputs of different networks is minimal and the surfaces largely converge. When the network is uncertain, the predictions of different models vary widely, producing a range of outputs, any of which could be the correct surface.
The spread in the predictions of a “committee of neural networks” is the “uncertainty” at that point. A good model should not just indicate the best prediction, but also indicates the uncertainty about each of these predictions. It’s like the neural network says “this property for material A will have a value of X and I’m highly confident about it.”
This could have been an elegant solution but for the sheer scale of the combinatorial space. “Each simulation (which is ground feed for the neural network) may take from tens to thousands of CPU hours,” explains Schwalbe-Koda. For the results to be meaningful, multiple models must be run over a sufficient number of points in the PES, an extremely time-consuming process.
Instead, the new approach only samples data points from regions of low prediction confidence, corresponding to specific geometries of a molecule. These molecules are then stretched or deformed slightly so that the uncertainty of the neural network committee is maximized. Additional data are computed for these molecules through simulations and then added to the initial training pool.
The neural networks are trained again, and a new set of uncertainties are calculated. This process is repeated until the uncertainty associated with various points on the surface becomes well-defined and cannot be decreased any further.
Gómez-Bombarelli explains, “We aspire to have a model that is perfect in the regions we care about (i.e., the ones that the simulation will visit) without having had to run the full ML simulation, by making sure that we make it very good in high-likelihood regions where it isn’t.”
The paper presents several examples of this approach, including predicting complex supramolecular interactions in zeolites. These materials are cavernous crystals that act as molecular sieves with high shape selectivity. They find applications in catalysis, gas separation, and ion exchange, among others.
Because performing simulations of large zeolite structures is very costly, the researchers show how their method can provide significant savings in computational simulations. They used more than 15,000 examples to train a neural network to predict the potential energy surfaces for these systems. Despite the large cost required to generate the dataset, the final results are mediocre, with only around 80 percent of the neural network-based simulations being successful. To improve the performance of the model using traditional active learning methods, the researchers calculated an additional 5,000 data points, which improved the performance of the neural network potentials to 92 percent.
However, when the adversarial approach is used to retrain the neural networks, the authors saw a performance jump to 97 percent using only 500 extra points. That’s a remarkable result, the researchers say, especially considering that each of these extra points takes hundreds of CPU hours.
This could be the most realistic method to probe the limits of models that researchers use to predict the behavior of materials and the progress of chemical reactions.
Accelerating geometry optimization in molecular simulation
More information:
Daniel Schwalbe-Koda et al, Differentiable sampling of molecular geometries with uncertainty-based adversarial attacks, Nature Communications (2021). DOI: 10.1038/s41467-021-25342-8
Provided by
Massachusetts Institute of Technology
This story is republished courtesy of MIT News (web.mit.edu/newsoffice/), a popular site that covers news about MIT research, innovation and teaching.
Citation:
Using adversarial attacks to refine molecular energy predictions (2021, September 2)
retrieved 2 September 2021
from https://techxplore.com/news/2021-09-adversarial-refine-molecular-energy.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.