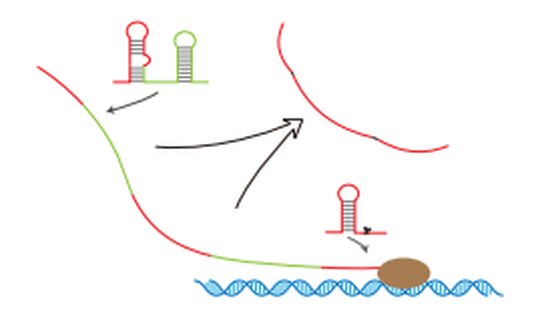
RNA is the central tenet of molecular biology, the stepping-stone between DNA and proteins.
In eukaryotes, messenger RNAs (mRNAs) undergo several processing steps (including splicing, and polyadenylation) to become functional mature mRNAs for producing proteins.
In this way mRNA processing plays a critical role during gene expression. This processing step, known as RNA maturation, happens inside the hub of the cell—the nucleus—where the mRNA has been freshly transcribed from DNA.
A question puzzling molecular biologists concerns how mRNA processing sites are precisely recognized; particularly how are they distinguished from surrounding sites with similar sequence content? There is currently a lack of knowledge of a general regulatory mechanism that recognizes actual sites during mRNA processing.
RNA structure, as an intrinsic characteristic of RNA molecules, was suggested to be involved in mRNA processing. However, how RNA structure contributes to the recognition of polyadenylation and splice sites, in general, remains elusive.
In a new study, researchers at the John Innes Centre have discovered the functional role of RNA structure in mRNA processing.
The team performed RNA structure profiling on Arabidopsis thaliana and generated the in vivo nuclear RNA structure landscape. This method allowed the dissection of pre-mRNA structure features before mRNA processing and determined the regulatory role of RNA structure during mRNA maturation.
Using a technique previously developed in the lab they captured the in vivo pre-mRNA structure over 12,000 genes with the nucleotide resolution and high accuracy.
The team found that nuclear mRNAs fold differently from cytosolic mRNAs. They identified distinctive pre-mRNA structure features which are responsible for both splicing and polyadenylation. The researchers further introduced mutations to disrupt these RNA structure features. They found that switching on or off these identified structure features is sufficient to regulate the splicing process.
The study, for the first time, unveils a new general RNA structure regulatory mechanism for mRNA processing. Additionally, the study emphasized the importance of dissecting RNA populations from different stages of the mRNA life cycle to investigate the relationship between RNA structure and biological functions.
This development in understanding how to regulate mRNA processing via RNA structure forms part of an emerging picture from the Ding lab of the importance of the shape of RNA in molecular biology.
Dr. Yiliang Ding, the corresponding author of the study said: “For the first time we understand the functional role of in vivo RNA structure during RNA maturation at a genome-wide scale. Because we understand the rules involved in this process, we could in future be able to engineer genes from a structural perspective rather than from a sequence perspective. We want to bring this insight to the attention of other researchers who can use it in plant and human health.”
In the future the team are hoping to ‘zoom out’ to explore the impacts of RNA structure on the growth and development we see in plants.
The study, “In vivo nuclear RNA structurome reveals RNA-structure regulation of mRNA processing in plants,” appears in Genome Biology.
mRNA is the stepping stone between DNA and proteins—the “M’ stands for messenger.
Between these three key stages of life (DNA to mRNA to protein) there are two processes: transcription and translation. In transcription DNA is transcribed to mRNA and then in translation mRNA is translated into protein. Together, these processes that describe the flow of genetic information are known as the central dogma of biology.
For mRNA to undergo translation and become a protein there is a processing stage, known as RNA maturation. In eukaryotes—organisms whose cells have a nucleus—his stage happens inside the nucleus, the hub of the cell—where the mRNA has been freshly transcribed.
The processing involves non-coding parts of the part mRNA being cut out and the parts which do code for proteins being stitched together to make a ‘clean’ instruction. This process is known as splicing.
The mRNA is then capped off at one end by what is known as a ‘Poly-A tail which marks the mRNA ready for export outside from the nucleus. Once this mature mRNA has left the nucleus, it enters the rest of the space inside the cell, called the cytoplasm, and here it is used as an instruction to build a protein through the translation process.
The underestimated mutation potential of retrogenes
More information:
Zhenshan Liu et al. In vivo nuclear RNA structurome reveals RNA-structure regulation of mRNA processing in plants, Genome Biology (2021). DOI: 10.1186/s13059-020-02236-4
Citation:
Study uncovers structural features regulating mRNA processing (2021, February 4)
retrieved 4 February 2021
from https://phys.org/news/2021-02-uncovers-features-mrna.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.