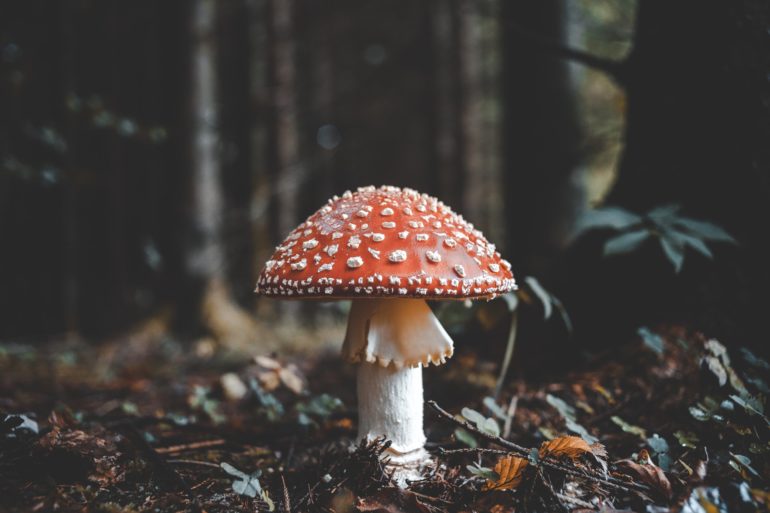
Though it might seem inanimate, the soil under our feet is very much alive. It’s filled with countless microorganisms actively breaking down organic matter, like fallen leaves and plants, and performing a host of other functions that maintain the natural balance of carbon and nutrients stored in the ground beneath us.
“Soil is mostly microorganisms, both alive and dead,” says Jennifer Bhatnagar, soil microbiologist and Boston University College of Arts & Sciences assistant professor of biology. It’s typical to see several hundred different types of fungi and bacteria in a single pinch of soil off the ground, she says, making it one of the most diverse ecosystems that exist.
Because there’s still so much unknown about soil organisms, until now scientists have not attempted to predict where certain species or groups of soil microbes live around the world. But having that knowledge about these organisms— too small to see with the naked eye—is key to better understanding the soil microbiome, which is made up of the communities of different microbes that live together.
A team of BU biologists, including Bhatnagar, took on that challenge—and their research reveals, for the first time, that it is possible to accurately predict the abundance of different species of soil microbes in different parts of the world. The team recently published their findings in a new paper in Nature Ecology & Evolution.
“If we know where organisms are on earth, and we know how they change through space and time due to different environmental forces, and something about what different species are doing, then we can much better predict how the function of these communities will change in terms of carbon and nutrient cycling,” Bhatnagar says. That kind of knowledge would have huge implications for agriculture, climate change, and public health.
“The health of the soils is so tied to the soil microbes,” says Michael Dietze, senior author on the study and a BU College of Arts & Sciences professor of earth and environment. Dietze, Bhatnagar, and researchers from their labs joined forces to work on this project, which involved analyzing hundreds of soil samples collected by National Ecological Observatory Network (NEON) research sites. Bhatnagar and her lab members brought to the team their soil expertise, while Dietze and his lab offered their unique ability to develop precise ecological forecasts and near-term environmental predictions.
The team learned that microbe predictability increases as spatial area increases, so the bigger the piece of land their model makes forecasts about, the more likely the predictions about what types of microbes live there will be accurate.
Dietze says the ability to accurately predict which microbes would likely be found in a given soil sample also increased as the researchers looked at organism groupings higher up on the phylogenetic scale, a system that classifies organisms based on evolutionary relatedness. On the smallest end of the scale, a “species” represents the finest level of classification; on the other end, a “phylum” makes up the largest and most diverse groupings of organisms. They were surprised to find that they were better able to predict the presence of a whole phylum, as opposed to individual species.
After receiving the genomic data of the soil samples from NEON, the research team’s forecasting models take into account environmental factors specific to the place where the soil came from—what plants live there, the soil acidity (pH), temperature, climate, and many others. They found their model was best able to predict the presence of microorganisms based on their symbiotic relationship with local plant species. Mycorrhizal fungi, for example, is a very common soil microbe that about 90 percent of plant families associate with, including pines and oak trees in New England.
In contrast, the team found it was more difficult to predict large groups of organisms based on their relationship with soil acidity. Despite knowing soil acidity levels, and what types of bacteria would typically like to live in that environment, their model couldn’t accurately predict the amount of bacteria that were actually present in the soil sample, Bhatnagar says. “That means there is something else beyond the relationship with [acidity], beyond the relationship with any other environmental factor that we typically measure in our ecosystems,” she says.
Now, Dietze and Bhatnagar’s team are expanding their forecasts beyond predicting microbes based on only their location, to also include specific times of the year.
“Building a framework for forecasting the soil microbiome at sites across the US will improve our understanding of seasonal and interannual change,” says Zoey Werbin, a Ph.D. student working in Bhatnagar’s lab and an author on the paper. “This could help us anticipate how climate change could affect microbial processes like decomposition or nitrogen cycling.”
With her dissertation project, Werbin hopes to answer fundamental questions about how and why the soil microbiome varies over time and space.
“The more we learn, the more we realize how important soil microbes are for agriculture, public health, and climate change. It’s really exciting to investigate how microscopic organisms can have such large-scale effects,” Werbin says. “We know certain factors, like temperature and moisture, affect microbial communities. But we don’t know how important those factors are compared to natural variability, or interactions between microbes. My Ph.D. project will help identify the driving forces of the soil microbiome, as well as the biggest sources of uncertainty.”
Soil bacteria evolve with climate change
More information:
Colin Averill et al, Soil microbiome predictability increases with spatial and taxonomic scale, Nature Ecology & Evolution (2021). DOI: 10.1038/s41559-021-01445-9
Citation:
Why scientists want to solve an underground mystery about where microbes live (2021, June 4)
retrieved 5 June 2021
from https://phys.org/news/2021-06-scientists-underground-mystery-microbes.html
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.